Computational nanoplasmonics in the quasistatic limit for biosensing applications
 Preprint: arXiv 1812.10722 Dec. 31, 2018.
Submitted: Dec. 31, 2018; Jan. 20, 2019; Mar. 20, 2019.
Accepted: Nov. 1. Published: Dec. 16, 2019.
DOI: 10.1103/PhysRevE.100.063305
Preprint: arXiv 1812.10722 Dec. 31, 2018.
Submitted: Dec. 31, 2018; Jan. 20, 2019; Mar. 20, 2019.
Accepted: Nov. 1. Published: Dec. 16, 2019.
DOI: 10.1103/PhysRevE.100.063305Abstract
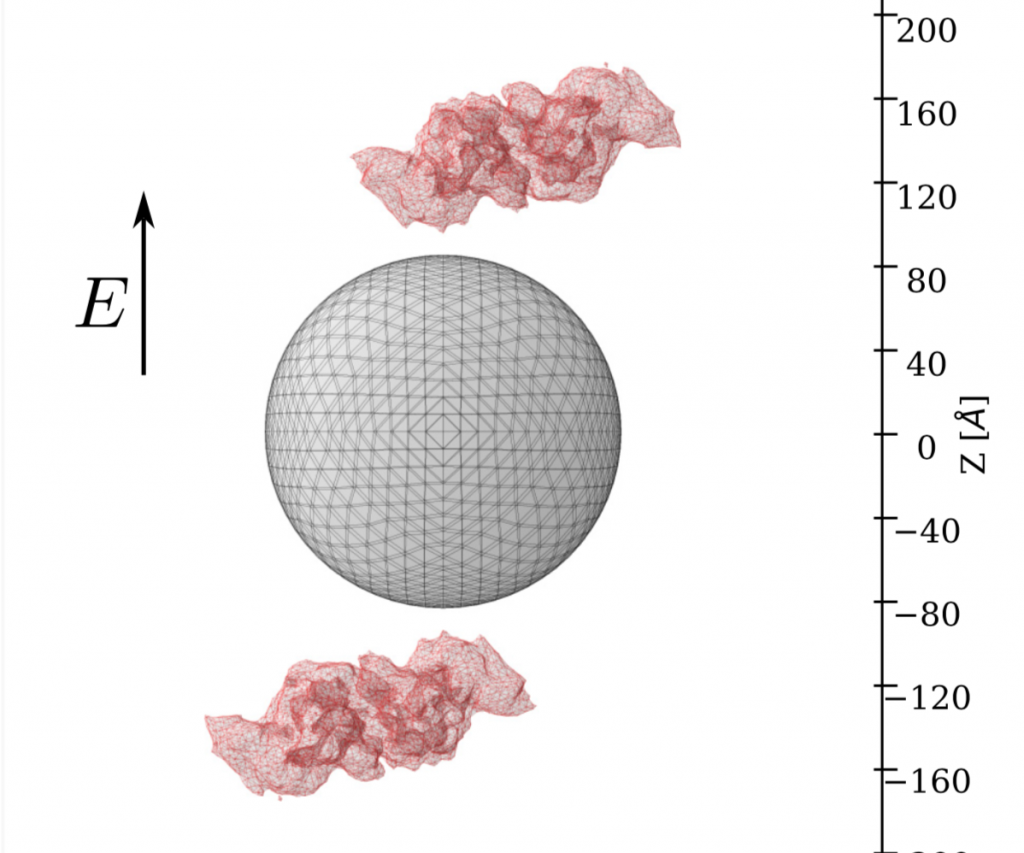
This work uses the long-wavelength limit to compute LSPR response of biosensors, expanding the open-source PyGBe code to compute the extinction cross-section of metallic nanoparticles in the presence of any target for sensing. The target molecule is represented by a surface mesh, based on its crystal structure. PyGBe is research software for continuum electrostatics, written in Python with computationally expensive parts accelerated on GPU hardware, via PyCUDA. It is also accelerated algorithmically via a treecode that offers O(N log N) computational complexity. These features allow PyGBe to handle problems with half a million boundary elements or more. Using a model problem consisting of an isolated silver nanosphere in an electric field, our results show grid convergence as 1/N, and accurate computation of the extinction cross-section as a function of wavelength (compared with an analytical solution). For a model of a sensor-analyte system, consisting of a spherical silver nanoparticle and a set of bovine serum albumin (BSA) proteins, our results again obtain grid convergence as 1/N (with respect to the Richardson extrapolated value). Computing the LSPR response as a function of wavelength in the presence of BSA proteins captures a red-shift of 0.5 nm in the resonance frequency due to the presence of the analytes at 1-nm distance. The final result is a sensitivity study of the biosensor model, obtaining the shift in resonance frequency for various distances between the proteins and the nanoparticle. All results in this paper are fully reproducible, and we have deposited in archival data repositories all the materials needed to run the computations again and re-create the figures. PyGBe is open source under a permissive license and openly developed. Documentation is available at http://barbagroup.github.io/pygbe/docs/
Reproducibility Packages
Documentation of the results presented in the paper, manuscript source files (.tex), Docker files, and supplementary materials are shared in the paper's GitHub repository: https://github.com/barbagroup/pygbe_lspr_paper
Reproducibility packages for several figures in the paper are also available on Figshare, at https://doi.org/10.6084/m9.figshare.c.4346156
Reference
- "Computational nanoplasmonics in the quasistatic limit for biosensing applications", Natalia C. Clementi, Christopher D. Cooper, Lorena A. Barba. Physical Review E, 100:063305 (Dec. 2019). 10.1103/PhysRevE.100.063305 // Preprint arXiv:1812.10722 // paper repository // reproducibility packages
News
- A figure from this paper was featured in the Physical Review E Kaleidoscope for December 2019