High-productivity, high-performance workflow for virus-scale electrostatic simulations with Bempp-Exafmm
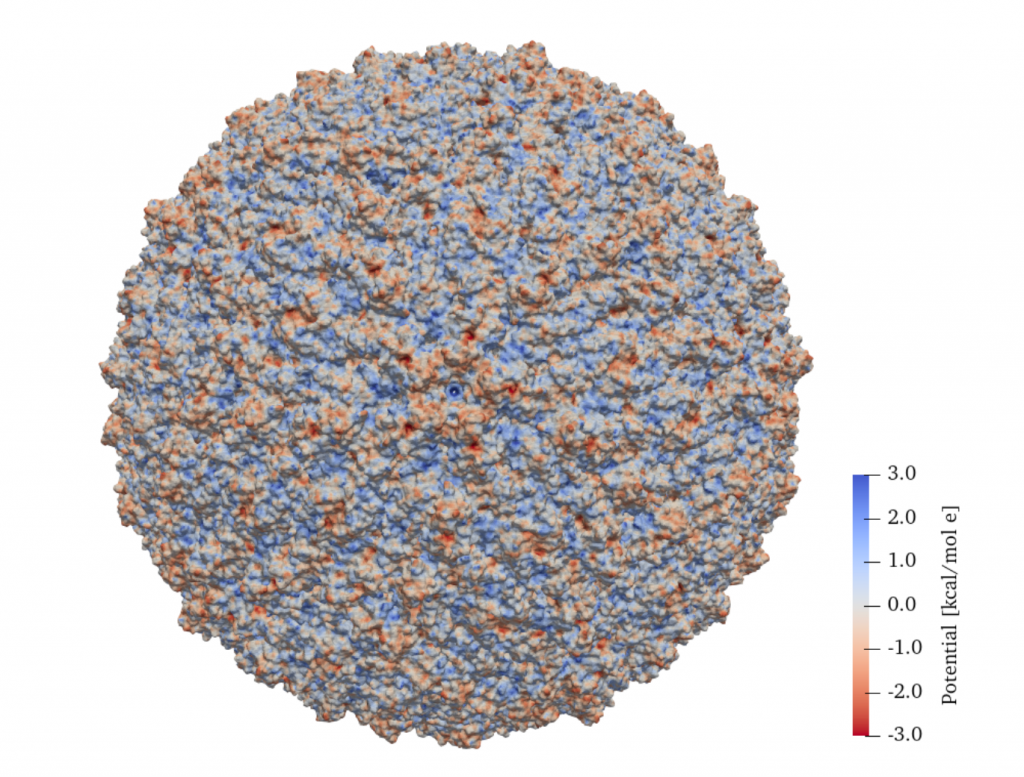
Result from our paper, showing the surface electrostatic potential on a Zika virus.
Preprint: arXiv:2103.01048, 1 Mar 2021. Submitted: 22 Mar. 2021.Abstract
Biomolecular electrostatics is key in protein function and the chemical processes affecting it.Implicit-solvent models expressed by the Poisson-Boltzmann (PB) equation can provide insights with less computational power than full atomistic models, making large-system studies—at the scale of viruses, for example —accessible to more researchers. This paper presents a high-productivity and high-performance computational workflow combining Exafmm, a fast multipole method (FMM) library, and Bempp, a Galerkin boundary element method (BEM) package. It integrates an easy-to-use Python interface with well-optimized computational kernels that are written in compiled languages. Researchers can run PB simulations interactively via Jupyter notebooks, enabling faster prototyping and analyzing. We provide results that showcase the capability of the software, confirm correctness, and evaluate its performance with problem sizes between 8,000 and 2 million boundary elements. A study comparing two variants of the boundary integral formulation in regards to algebraic conditioning showcases the power of this interactive computing platform to give useful answers with just a few lines of code. As a form of solution verification, mesh refinement studies with a spherical geometry as well as with a real biological structure (5PTI) confirm convergence at the expected 1/N rate, for N boundary elements. Performance results include timings, breakdowns, and computational complexity. Exafmm offers evaluation speeds of just a few seconds for tens of millions of points, and O(N) scaling. This allowed computing the solvation free energy of a Zika virus, represented by 1.6 million atoms and 10 million boundary elements, at 80-min runtime on a single compute node (dual 20-core Intel Xeon Gold 6148). All results in the paper are presented with utmost care for reproducibility. Readers can find input data (meshes and pqr files) in a Zenodo archive, Jupyter notebooks and secondary data necessary to reproduce all the figures in the paper's GitHub repository at https://github.com/barbagroup/bempp_exafmm_paper/, and all the software in version-controlled repositories under permissive standard licenses.
References
- "High-productivity, high-performance workflow for virus-scale electrostatic simulations with Bempp-Exafmm (manuscript and repro-pack)", Tingyu Wang, Christopher D. Cooper, Timo Betcke, Lorena A. Barba. (2021). 10.5281/zenodo.4568951
Archive of the manuscript GitHub repository, upon submission of the paper.
- "High-productivity, high-performance workflow for virus-scale electrostatic simulations with Bempp-Exafmm (meshes and pqr files)", Tingyu Wang, Christopher D. Cooper, Timo Betcke, Lorena A. Barba. (2021). 10.5281/zenodo.4568768
Reproducibility package.
- "High-productivity, high-performance workflow for virus-scale electrostatic simulations with Bempp-Exafmm", Tingyu Wang, Christopher D. Cooper, Timo Betcke, Lorena A. Barba. (March 2021). Preprint arXiv:2103.01048 // manuscript repository // archive: meshes and pqr files // archive: manuscript and repro-pack
On Twitter
https://twitter.com/LorenaABarba/status/1376501893564833793